1H-13C
HMQC-TOCSY
The 1H-13C HMQC-TOCSY experiment combines 1H-13C HMQC and 1H-TOCSY
experiments to give through-bond correlations between a 13C-attached 1H to all
other coupled 1H. The coupled 1H's can be seen along a line at the same 13C
chemical shift from the carbon atom attached to the primary 1H. The spectrum
should be compared with a 1H-13C HSQC or HMQC spectrum. The differences between
a HSQC-TOCSY and HMQC-TOCSY is the same as mentioned
here. The HSQC-TOCSY sequence is generally recommended over the HMQC
version.
The following figure is the Varian 2D gradient-selected, phase-sensitive
1H-13C HMQC-TOCSY pulse sequence (varian name: gHMQCTOXY) with
the main parameters displayed (NMR500).
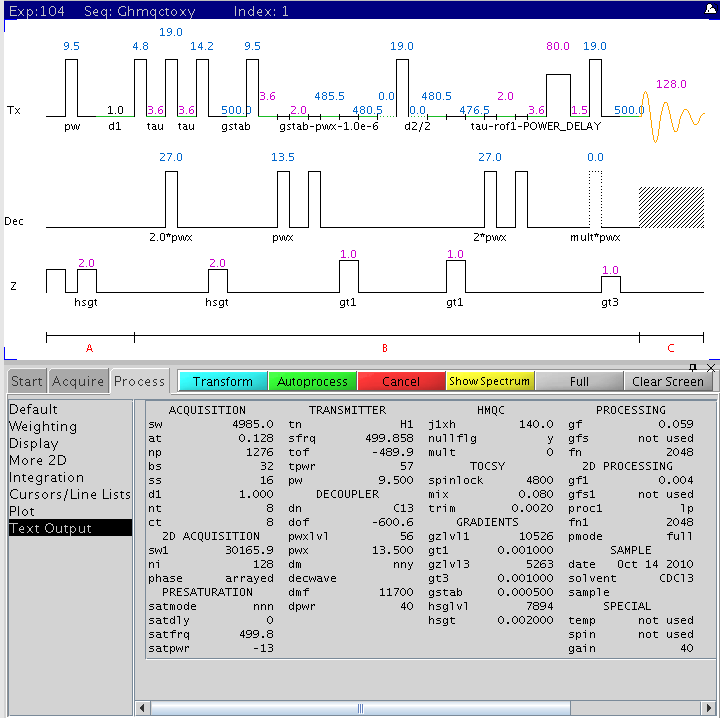
Procedure (on NMR500)
- Temperature must be regulated.
- Turn spin OFF.
- Gradients amplifier should be turned on (displays
RUN).
- Type pfgon='nny' su to allow gradient pulses.
- Lock and shim your sample as usual. Keep lock at ~80% after
shimming.
- Find proper sw and tof to use
- Under exp1 (type jexp1 to go
there), collect a standard 1H spectrum (nt=1 is Ok for this, but 4 gives
cleaner spectrum)
- Put the box cursor to enclose the signal region
and to include ~10% flat baseline area at each end of the peak region. Type
movesw. This command sets a new spectral width
sw and spectrum center tof.
- Re-collect a 1D 1H spectrum with the new setting.
Type wft f full. Reference the 1H spectrum carefully.
- Load standard 2D ghmqctoxy data file
- Join a different experiment (i.e.
jexp2 or create it with cexp(2) if it
doesn't exist).
- From exp2, type mf(1,2) to move
fid from exp1 to exp2. Type wft f full to display
spectrum. Check reference.
- Type setexp('ghmqctoxy')
su to load standard parameters. Solvent, 1H spectral
width, and center are copied over from 1D spectrum.
- Tune 1H and X channels. Accuracy of pre-calibrated pw
depends on probe tuning.
- Recable the X nucleus probe input cable.
- Disconnect the X nucleus cable at the filters in
front of the broadband preamp (left side). Connect the X-channel probe
input cable to one end of the long, 13C bandpass filter (labeled 13C on
filter) on the floor. Connect the other end of the filter to the X-channel
decoupler cable coming from the back of the magnet leg interface (long,
tall box to the right of the magnet).
- Re-shim Z1 and Z2 a bit after this cabling
change.
- Check/set proper 1H transmitter power and pulse width. For cdcl3 or other
solvents with similar dielectric properties, use the default. Otherwise,
calibrate pw according to this procedure.
- Use default parameters values (~40 mins experiment) except the
following:
- pw=XXX pw90=pw
(value calibrated with your solvent at tpwr=57)
- Optionally, for better quality data (may results
in longer data collection time):
- ss=32 (more dummy scans before
data collection)
- nt=16 (or 24, 32, ... 8*n) for
more sensitivity and less artifacts
- ni=256 (better resolution along
indirect dimension)
- Type time to check experiment time.
- Type go to collect data.
- Watch for ADC or receiver overflow:
- In arrayed experiments such as 2D's, a fixed
gain (not autogain) must be set. Watch the receiver overflow light and the
ADC overflow error message immediately after the experiment starts. If the
receiver overflows (red light blinks on VT display) or ADC overflow message
is seen above the command area anytime during the experiment except during
the dummy scan period, it is likely the data are ruined. Stop the
experiment and reduce gain by 2-4 at a time until the overflow disappears.
The default (gain=36) works most of the time unless there
is an extremely strong 1H signal in your sample. With a gain above 20, it
makes only small differences.
- After the experiment is done, recable the X (decoupler) channel
connection to the original configuration.
Default parameters
The default setting (modified from Varian's default values) in our parameter
set uses the following for a quick 2D experiment:
- ni=128 (128 complex points along indirect dimension. For
better resolution, set ni bigger, ie.
ni=256)
- d1=1 (recycle delay between scans is 1 sec)
- ss=16 (16 dummy scans before actual data collection)
- nt=8 (8 scans)
- phase=1,2 (phase-sensitive mode in indirect
detection)
- gain=40 (reduce it by 2 at a time if receiver or ADC
overflows)
- sspul='y' (spin randomization before recycle delay with
homospoil gradients turned on)
- mix=0.080 (TOCSY spinlock mixing time is 80 msec)
Data Processing
You can use vnmrJ to process the data by clicking
autoprocess, but vnmrJ's graphics display for 2D spectrum
leaves much to be desired. A separate program is strongly recommended for
detailed processing and analysis.
One of the most widely used free processing/display software package for
multi-dimensional NMR data processing is nmrPipe/nmrDraw. This
program package has been installed on NMR500 and the data station. Follow this procedure to process data with nmrPipe/nmrDraw.
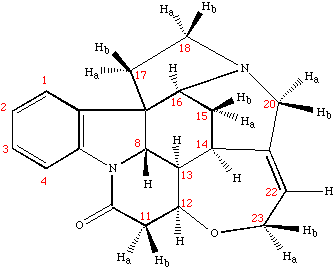
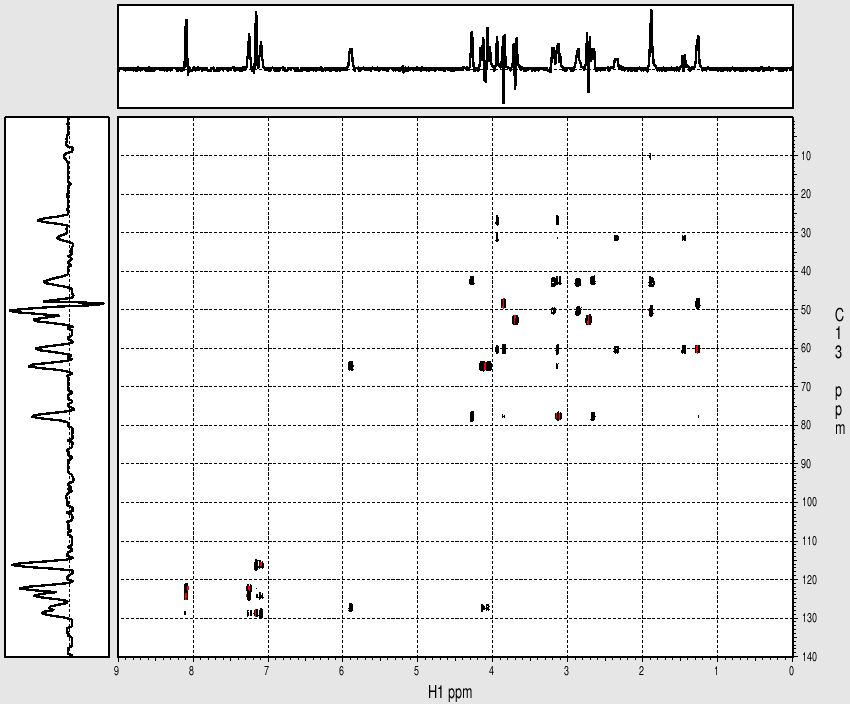
Example spectrum collected on NMR500
Sample: Strychnine at ~ 25mg/mL (~ 100mM) in cdcl3
- Data collected October 2010 (~43 mins with nt=8)
- TOCSY mixing time: 80 msec
- Processed and displayed with nmrPipe/nmrDraw
|
 |
H. Zhou
Updated Nov 2010



